Context
The composition and concentration of bioaerosols in the air is a critical parameter to consider for the welfare of animals and humans. Monitoring bioaerosols by air sampling is one of the most efficient methods available. Nonetheless, the detection of DNA from bioaerosols can be challenging without the appropriate tools and the optimization of sampling and detection protocols. The use of the Coriolis µ air sampler itself is fairly simple thanks to its user-friendly design and cyclonic collection on liquid. Nonetheless, there are many other factors that need to be considered during the workflow in order to obtain the best results possible.
In this Application Note we provide, through examples, a guide on How to analyze your Coriolis µ air sample for the detection of bacterial DNA. The recommendations of this guide will allow you to verify the effectiveness of your air sampling protocol, by confirming that you have successfully collected bacterial DNA from the air.
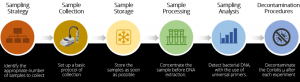
Sampling strategy: Choose a sampling strategy adapted to the environment (room size, air flow patterns, etc). Place the Coriolis µ at least 1 meter above ground level. Depending on the size of the sampling area, plan to collect at several different points across the area, if possible.
Sample collection: As a basis for validating the technique for DNA extraction, set up a basic protocol of collection of 300L/min for 10 minutes using 15 ml of collection liquid, either indoors or outdoors. Then, based on the results obtained, adjust the collection time.
Sample storage: For the detection of DNA, we recommend to store the samples as soon as possible at 4 °C or -20 °C, for long term storage.
Sample processing: To increase the yield of DNA, given the relatively low concentration of bacteria expected in a sample of 1 m3 of air, we highly recommend to concentrate the sample before DNA extraction. For sample concentration, use either a filtration (Amicon Ultra 15 100 Kda) or a centrifugation (>16000 rpm for 10 minutes) technique.
Sample Analysis
DNA extraction: The extraction of bacterial DNA can be carried out with the technique of your choice. We recommend the use of a kit specific for bacterial DNA.
qPCR: For the detection of bacterial DNA, the use of universal primers for the amplification of 16S rRNA is a current practice in molecular biology. For this reason, it is also important to select a mix that is sensitive and precise enough to perform reliable PCR assays.
Decontamination: Decontaminate the Coriolis µ after each experiment. The cane and the air intake can be autoclaved.